Publishing
Data produced by BLISS sessions is published into Redis (RAM storage).
In Redis, data is stored for a limited period of time (1 day by default) and for a limited amount (1GB by default).
Experiment and data files structure¶
A complete experiment can be seen has a succession of measurements with different samples in different conditions.
In a Bliss session, the experiment can be seen as a tree, where the trunk is the session itself and where each measurement performed for a given sample and experimental conditions is a branch.
As an example, let’s consider:
- two samples: ‘sample1‘ and ‘sample2‘
- a session named ‘test_session‘
- several measurements with the two samples
- a measurement that consists in scanning one of the samples by moving a motor ‘roby‘ and measuring the value of a counter ‘diode‘ along the scan.
The sample scan is performed with the following command (for details, see: scan commands):
ascan(roby, 0, 9, 10, 0.1, diode)
- To start the session:
bliss -s test_session
- Format data saving path for the experiment:
While starting a session for the first time, a directory with the same name as the session is created and the scan data is stored in this directory.
As two different samples will be scanned, one sub-directory per sample will be created. To do that, the SCAN_SAVING object has to be used. The data saving path is customized by adding a new parameter ‘s_name‘ usable in the template of the PATH.
SCAN_SAVING.add('s_name', '') # add a parameter named "s_name".
SCAN_SAVING.template = '{session}/{s_name}/' # modify data saving path template.
SCAN_SAVING.s_name = 'sample1' # set value of parameter "s_name".
- Perform a first measurement:
ascan(roby, 0, 9, 10, 0.1, diode)
- Perform a second measurement:
ascan(roby, 0, 9, 10, 0.1, diode)
- Change the data saving path for measurements on sample2:
SCAN_SAVING.s_name = 'sample2'
- Perform a measurement:
ascan(roby, 0, 9, 10, 0.1, diode)
For this experiment the files structure inside the session main folder is described by the following tree:
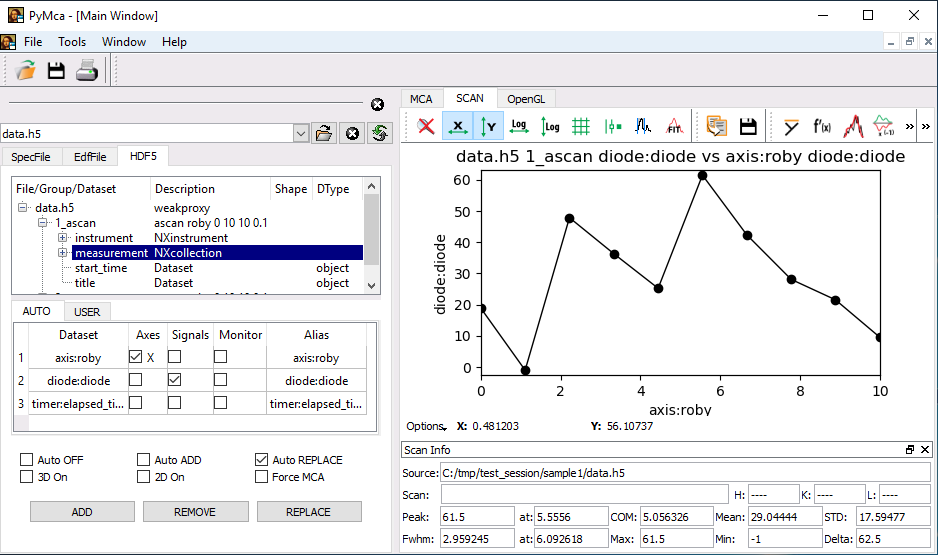
The measurements data can be accessed by reading the content of the hdf5 files (see silx, pymca). One file usually contains multiple scans data.